Microbial mediators of food web dynamics
Exploring bacteria-eukaryote interactions in the Great Lakes
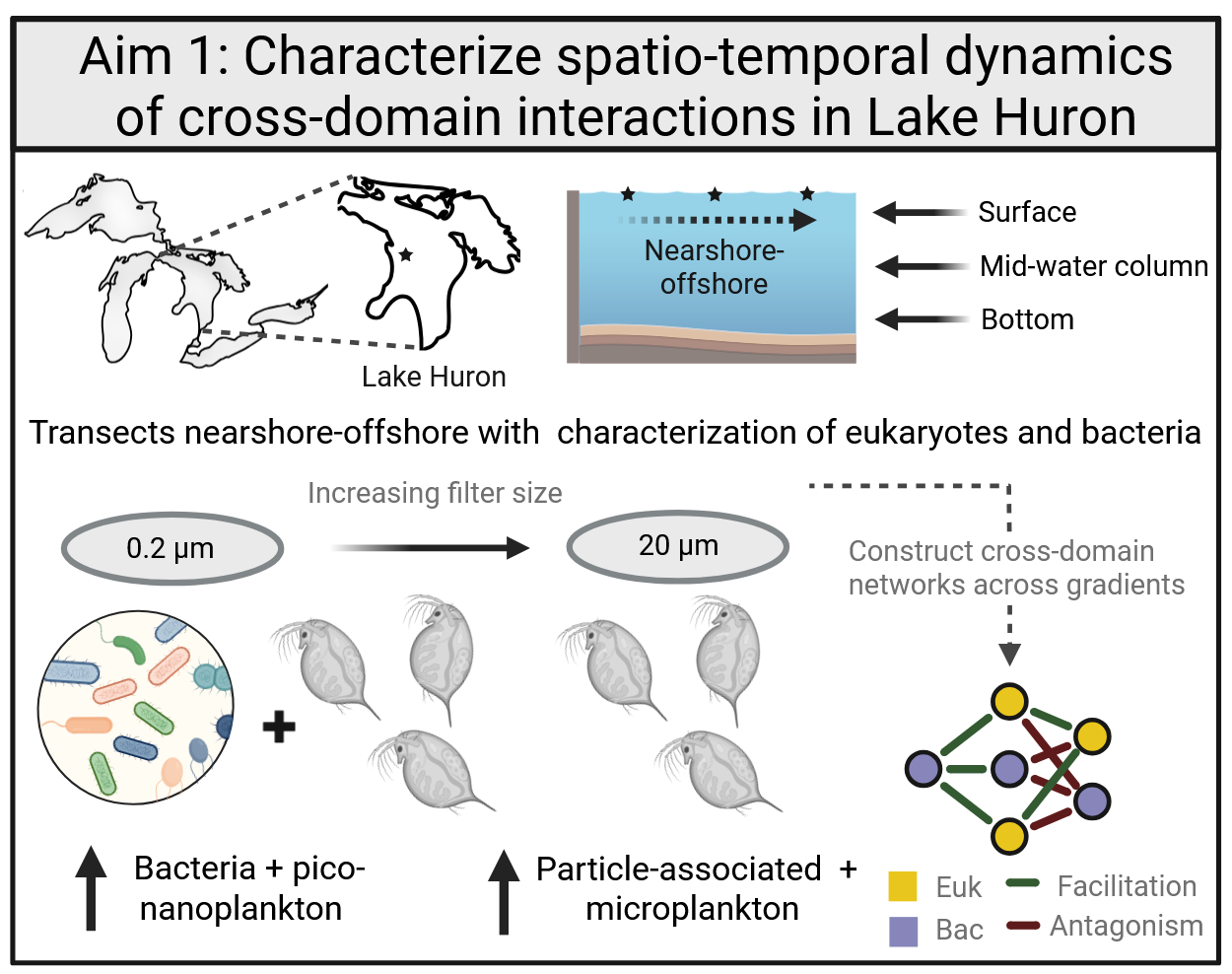
In my current position I work with Cooperative Institute for Great Lakes Research on several projects in collaboration with the Michigan Geomicrobiology Laboratory. Two on-going projects aim to characterize bacterial-eukaryotic interactions in the Great Lakes, especially in the context of Dreissenid mussel invasion (i.e., quagga mussels and zebra mussels) and Microcystis dominated cyanobacterial harmful algal blooms (cHABs). These projects leverage paired amplicon sequencing data of the 16S and 18S rRNA genes in Lake Huron and timeseries shotgun metagenomics data from the western basin of Lake Erie, respectively. Fundamental shifts in the flow of energy are expected under high size-selective filtering and phosphorus depletion mediated by Dreissenid mussels, potentially resulting in decoupling of primary productivity by phytoplankton and heterotrophic zooplankton grazing. I am exploring the possibility of compensatory dietary and/or interactive shifts under high mussel grazing regimes in oligotrophic (low nutrient) areas of Lake Huron as indicated by changes in network-inferred interactions between organisms.
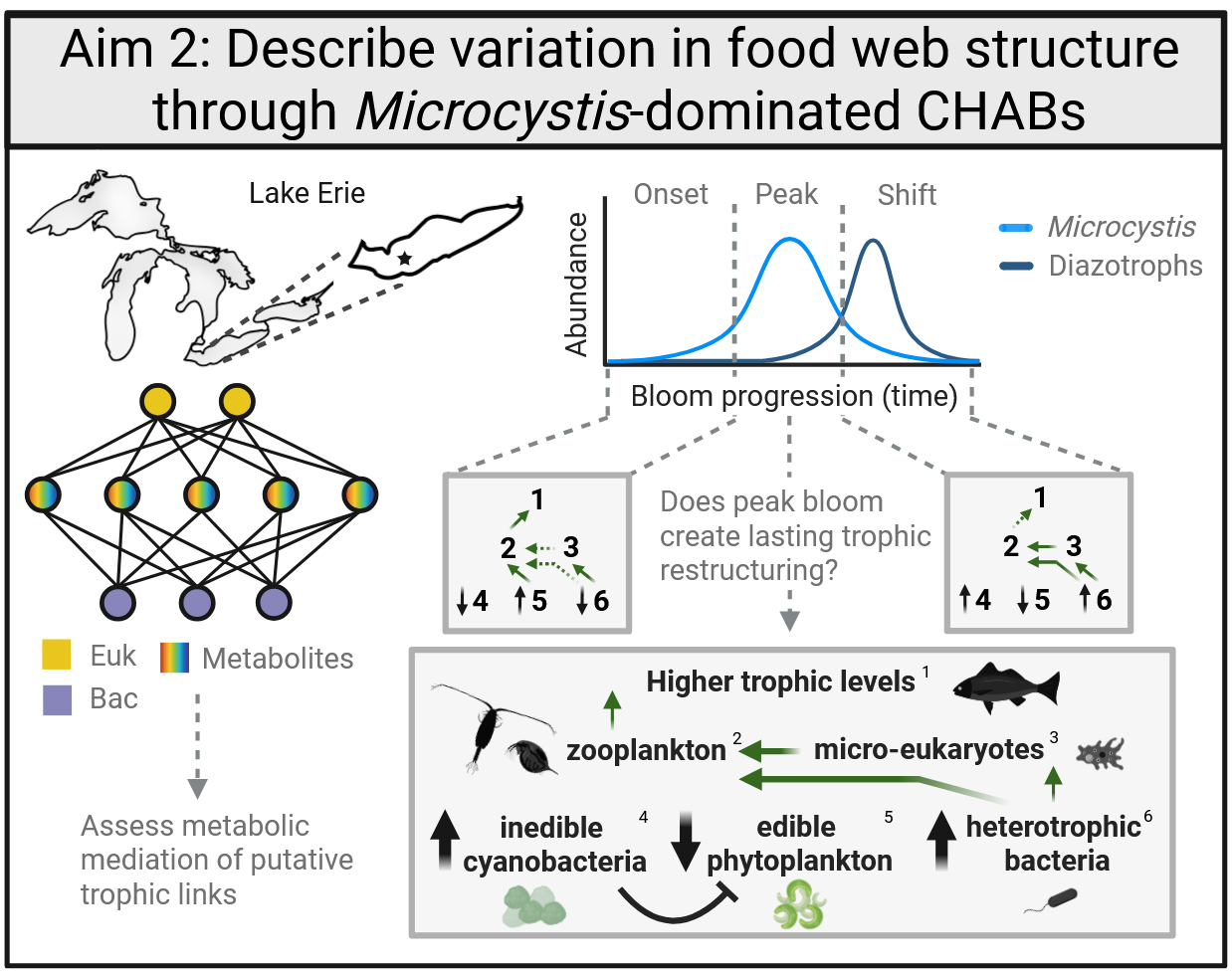
Where Lake Huron is fairly nutrient-poor these days, the western basin of Lake Erie is much more nutrient rich, experiecing seasonally recurrent cHABs. Microcystis is a cyanobacterium that commonly dominates these blooms, taking advantgae of high nitrogen and phosphorus to outcompete other phytoplankton taxa. These colonial cyanos also produce harmful compounds which may modulate interactions with other bacterioplankton and/or eukaryotes. Shotgun metagenomics over time allows us to explore all the DNA in a sample in a temporally-resolved manner. Paired with metabolomes, I am hoping to construct tripartite networks to understand community interacitions within and between bacteria and eukaryotes as informed by metabolism and environmental chemistry.